 Retroviruses & Structural Biochemistry
Retroviruses & Structural Biochemistry
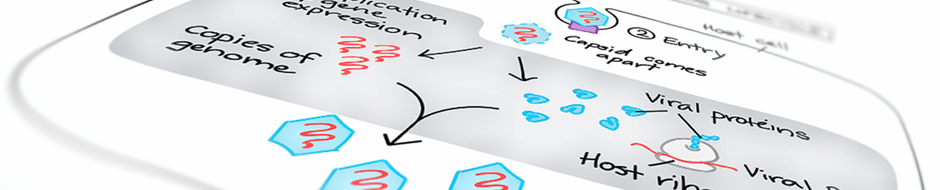
 Retroviruses & Structural Biochemistry
Retroviruses & Structural Biochemistry

 are characterized by their ability to gain resistance from therapeutic or vaccinal intervention through the accumulation of escape mutations in the viral genome and/or the interaction of regulatory viral proteins with cellular pathways. Understanding the structural determinants of the functions of the viral proteins is crucial for the development of efficient antiviral therapies.
are characterized by their ability to gain resistance from therapeutic or vaccinal intervention through the accumulation of escape mutations in the viral genome and/or the interaction of regulatory viral proteins with cellular pathways. Understanding the structural determinants of the functions of the viral proteins is crucial for the development of efficient antiviral therapies.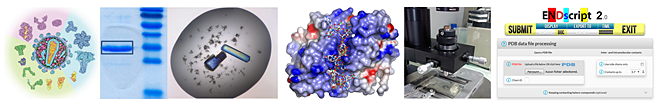
 ), observed a new dimeric form of an avian integrase domain (Ballandras et al., 2011
), observed a new dimeric form of an avian integrase domain (Ballandras et al., 2011  ) and determined the crystal structures of the matrix and capsid proteins of the Feline Immunodeficiency Virus (FIV) (Serrière et al., 2013
) and determined the crystal structures of the matrix and capsid proteins of the Feline Immunodeficiency Virus (FIV) (Serrière et al., 2013  ; Folio et al., 2017
; Folio et al., 2017  ).
).
 , in coll. with O. Marcillat), the bacterial type 2 secretion system (Pineau et al., 2014
, in coll. with O. Marcillat), the bacterial type 2 secretion system (Pineau et al., 2014  , in coll. with V. Shevchik) and the Ser/Thr kinase StkP of S. pneumoniae (Righino et al., 2017
, in coll. with V. Shevchik) and the Ser/Thr kinase StkP of S. pneumoniae (Righino et al., 2017  , in coll. with MC. de Rosa; Zucchini et al., 2018
, in coll. with MC. de Rosa; Zucchini et al., 2018  , in coll. with C. Grangeasse).
, in coll. with C. Grangeasse). / ENDscript
/ ENDscript  for protein sequence and structure analyses, which counts more than 15,000 users per year (Robert & Gouet, 2014
for protein sequence and structure analyses, which counts more than 15,000 users per year (Robert & Gouet, 2014  ).
).
 By clicking on the PDB logo, you will be redirected to the Protein Data Bank website presenting the extensive list of the structures we solved @IBCP.
By clicking on the PDB logo, you will be redirected to the Protein Data Bank website presenting the extensive list of the structures we solved @IBCP. |

Back to top
| Deciphering key features in protein structures with the new ENDscript server |
|---|
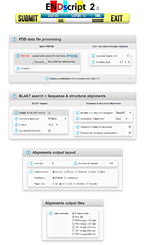 ENDscript 2 is a friendly webserver for extracting and rendering a comprehensive analysis of primary to quaternary protein structure information in an automated way. This major upgrade has been fully re-engineered to enhance speed, accuracy and usability with interactive 3D visualization. It takes advantage of the new version 3 of ESPript, our well-known sequence alignment renderer, improved to handle a large number of data with reduced computation time. From a single PDB entry or file, ENDscript produces high quality figures displaying multiple sequence alignment of proteins homologous to the query, colored according to residue conservation. Furthermore, the experimental secondary structure elements and a detailed set of relevant biophysical and structural data are depicted. All this information and more are now mapped on interactive 3D PyMOL representations. Thanks to its adaptive and rigorous algorithm, beginner to expert users can modify settings to fine-tune ENDscript to their needs. ENDscript has also been upgraded as an open platform for the visualization of multiple biochemical and structural data coming from external biotool webservers, with both 2D and 3D representations. ENDscript 2 and ESPript 3 are freely available at https://endscript.ibcp.fr
ENDscript 2 is a friendly webserver for extracting and rendering a comprehensive analysis of primary to quaternary protein structure information in an automated way. This major upgrade has been fully re-engineered to enhance speed, accuracy and usability with interactive 3D visualization. It takes advantage of the new version 3 of ESPript, our well-known sequence alignment renderer, improved to handle a large number of data with reduced computation time. From a single PDB entry or file, ENDscript produces high quality figures displaying multiple sequence alignment of proteins homologous to the query, colored according to residue conservation. Furthermore, the experimental secondary structure elements and a detailed set of relevant biophysical and structural data are depicted. All this information and more are now mapped on interactive 3D PyMOL representations. Thanks to its adaptive and rigorous algorithm, beginner to expert users can modify settings to fine-tune ENDscript to their needs. ENDscript has also been upgraded as an open platform for the visualization of multiple biochemical and structural data coming from external biotool webservers, with both 2D and 3D representations. ENDscript 2 and ESPript 3 are freely available at https://endscript.ibcp.fr  and https://espript.ibcp.fr and https://espript.ibcp.fr  . . Robert X & Gouet P (2014). Nucleic Acids Res. 42:W320-4 
|

Back to top
| Internal IBCP |
|---|
|
Vincent ChaptalMMSB, CNRS – Université de Lyon Christophe GrangeasseMMSB, CNRS – Université de Lyon Guillaume LaunayMMSB, CNRS – Université de Lyon Juliette MartinMMSB, CNRS – Université de Lyon Cédric OrelleMMSB, CNRS – Université de Lyon Bernard VerrierLBTI, CNRS – Université de Lyon |
| External |
|---|
|
Guzmán Álvarez TouronUniversidad de la República - Paysandú, Uruguay Yannick BlanchardViral Genetics and Biosafety - ANSES Ploufragan, France Laure GuyÉcole Normale Supérieure - Lyon, France Xavier HanoulleNMR & Molecular Interactions - CNRS Villeneuve d'Ascq, France Vincent ParissiFundamental Microbiology and Pathogenicity Lab - CNRS Bordeaux, France Stéphane PaulGroupe sur l'Immunité des Muqueuses et Agents Pathogènes - Faculté de Médecine de Saint-Etienne, France Corinne RonfortPathogenicity and Virus Vaccine Lab - INRA Grenoble, France Maria-Cristina de RosaInstitute of Chemistry of Molecular Recognition - Rome, Italy Guy SchoehnInstitute of Structural Biology - Grenoble, France Vladimir ShevchikMicrobiology, Adaptation and Pathogenesis Lab - INSA Villeurbanne, France |

Back to top

Back to top

 Protéines, 7 passage du Vercors,
Protéines, 7 passage du Vercors,